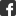Rätsel um Erkennung ungefalteter Proteine gelöst:
Das Schloss formt den Schlüssel
14.02.2011, Pressemitteilungen
Proteine erkennen sich normalerweise an ihrer spezifischen dreidimensionalen Struktur. Passt der Schlüssel zum Schloss, kann eine Reaktion stattfinden. Doch es gibt auch Reaktionen, bei denen der Schlüssel zu Beginn der Reaktion noch gar keine Form besitzt. Chemiker der Technischen Universität München (TUM) und der Max-Planck-Forschungsstelle für die Enzymologie der Proteinfaltung (Halle/Saale) haben nun an einem Beispiel gezeigt, wie das funktionieren kann. Ihre Ergebnisse erscheinen in dieser Woche in den Proceedings of the National Academy of Science (PNAS).
Wechselwirkungen zwischen Proteinen sind von grundlegender Bedeutung für eine Vielzahl von Prozessen in jeder lebenden Zelle. Doch um eine biologische Funktion ausüben zu können, müssen die Proteine erst ihre spezifische, dreidimensionale Form annehmen. In den letzten Jahren wurde eine Reihe von Reaktionen beschrieben, bei denen einer der Wechselwirkungspartner seine biologisch aktive Struktur erst während der Bindung einnimmt. Ein großes Rätsel blieb dabei, wie ihre Bindungspartner solche unstrukturierten Proteine überhaupt erkennen können.
Die Wissenschaftler um Professor Thomas Kiefhaber (TUM) stellten sich die Frage, ob für die Erkennung lokale Eigenschaften genügen oder ob der unstrukturierte Bindungspartner zunächst eine spezifische räumliche Struktur einnehmen muss. Dafür kämen regelmäßige Strukturelemente wie die wendeltreppenförmigen α-Helices oder β-Faltblätter in Frage, bei denen interne Wasserstoffbrückenbindungen ausgebildet werden.
In Zusammenarbeit mit der Gruppe von Professor Gunter Fischer (Max Planck Forschungsstelle für die Enzymologie der Proteinfaltung Halle/Saale) entwickelten die Wissenschaftler eine neue Methode, die es erstmals erlaubt, die Ausbildung einzelner Wasserstoffbrückenbindungen im Verlaufe eines Bindungsprozesses zu verfolgen.
Als Modellsystem benutzten sie das Enzym Ribonuklease S, das in seiner aktiven Form aus dem S-Protein und dem α-helikalen S-Peptid besteht. Während das S-Protein eine definierte dreidimensionale Form besitzt, ist das S-Peptid allein zunächst ungefaltet. Die Forscher untersuchten nun, ob das S-Protein das unstrukturierte S-Peptid oder eine geringe Population des Peptids in helikaler Konformation erkennt. Dafür wurden mit Hilfe chemischer Proteinsynthese gezielt Sauerstoffatome in Peptidbindungen des S-Peptids durch Schwefelatome ersetzt, wodurch einzelne Wasserstoffbrückenbindungen destabilisiert werden.
Zeitabhängige Messungen des Bindeprozesses der veränderten Peptide zeigten nun, dass sich die Wasserstoffbrücken im S-Peptid und damit die α-helikale Struktur erst nach der Bindung an das S-Protein ausbilden und somit für die Erkennung nicht wichtig sein können. Die Protein-Protein Erkennung hingegen erfolgt stattdessen über lokale hydrophobe Wechselwirkungen des S-Proteins mit zwei räumlich eng begrenzten Bereichen des unstrukturierten S-Peptids.
Diese Ergebnisse sind von grundlegender Bedeutung für das Verständnis des Mechanismus von Protein-Protein Wechselwirkungen. Die entwickelte Methode kann in Zukunft auch in anderen Systemen eingesetzt werden, um die Strukturbildung in Proteinen detailliert zu untersuchen.
Die Forschungsarbeit wurde unterstützt aus Mitteln des Bundesministeriums für Bildung und Forschung (ProNet-T3) und der Deutschen Forschungsgemeinschaft (Exzellenzcluster Munich Center for Integrated Protein Science).
Publikation:
Mapping backbone and side-chain interactions in the transition state of a coupled protein
folding and binding reaction, Annett Bachmann, Dirk Wildemann, Florian Praetorius, Gunter Fischer, and Thomas Kiefhaber
PNAS, Early Edition, Publikation Online in der Woche vom 14.02.2011, http://www.pnas.org/cgi/doi/10.1073/pnas.1012668108
Kontakt:
Prof. Dr. Thomas Kiefhaber
Technische Universität München
Lehrstuhl für Biophysikalische Chemie
Lichtenbergstr. 4, 85748 Garching, Germany
Tel: +49 89 289 13420 – Fax: +49 89 289 13416
E-Mail -
Internet
Kontakt: presse@tum.de
Mehr Information
| 110214_SideChain_DE_PW.pdf |
Druckversion der Presseinformation (DE),
(Type: application/pdf,
Größe: 105.8 kB)
Datei speichern
|
|
| 110215_SideChain_EN_PW.pdf |
Printable version of the press release (EN),
(Type: application/pdf,
Größe: 92.6 kB)
Datei speichern
|